Note
Go to the end to download the full example code or to run this example in your browser via Binder
Comparing initial sampling methods#
Holger Nahrstaedt 2020 Sigurd Carlsen October 2019
When doing baysian optimization we often want to reserve some of the early part of the optimization to pure exploration. By default the optimizer suggests purely random samples for the first n_initial_points (10 by default). The downside to this is that there is no guarantee that these samples are spread out evenly across all the dimensions.
Sampling methods as Latin hypercube, Sobol’, Halton and Hammersly take advantage of the fact that we know beforehand how many random points we want to sample. Then these points can be “spread out” in such a way that each dimension is explored.
See also the example on an integer space sphx_glr_auto_examples_initial_sampling_method_integer.py
def plot_searchspace(x, title):
fig, ax = plt.subplots()
plt.plot(np.array(x)[:, 0], np.array(x)[:, 1], 'bo', label='samples')
plt.plot(np.array(x)[:, 0], np.array(x)[:, 1], 'bo', markersize=80, alpha=0.5)
# ax.legend(loc="best", numpoints=1)
ax.set_xlabel("X1")
ax.set_xlim([-5, 10])
ax.set_ylabel("X2")
ax.set_ylim([0, 15])
plt.title(title)
n_samples = 10
space = Space([(-5.0, 10.0), (0.0, 15.0)])
# space.set_transformer("normalize")
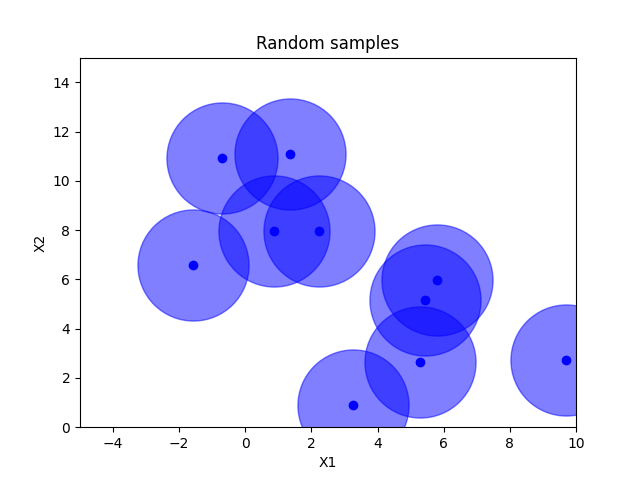
Random sampling#
x = space.rvs(n_samples)
plot_searchspace(x, "Random samples")
pdist_data = []
x_label = []
pdist_data.append(pdist(x).flatten())
x_label.append("random")
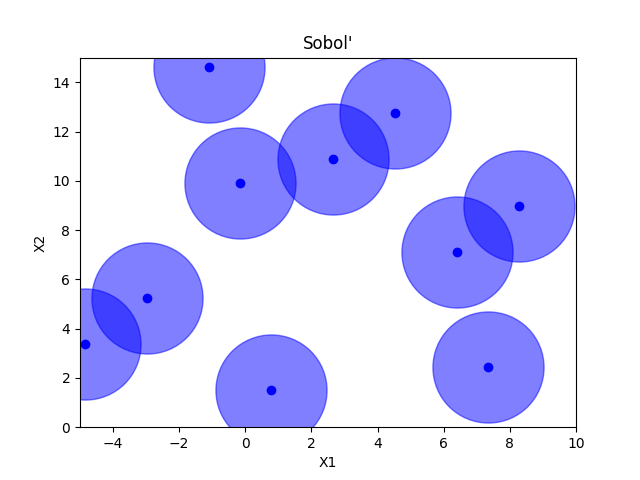
Sobol’#
D:\git\scikit-optimize\skopt\sampler\sobol.py:521: UserWarning: The balance properties of Sobol' points require n to be a power of 2. 0 points have been previously generated, then: n=0+10=10.
warnings.warn(
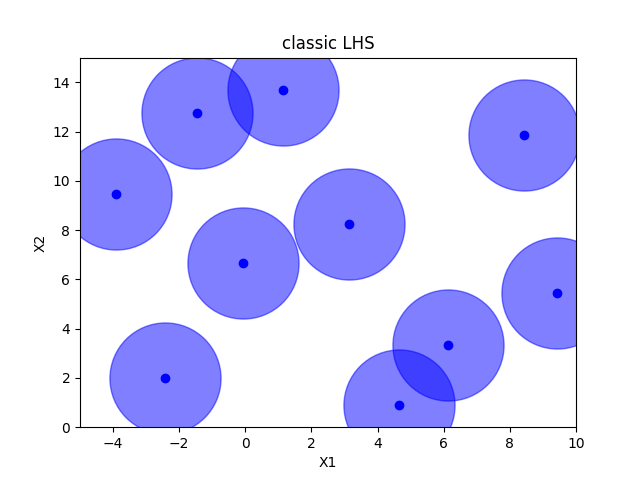
Classic Latin hypercube sampling#
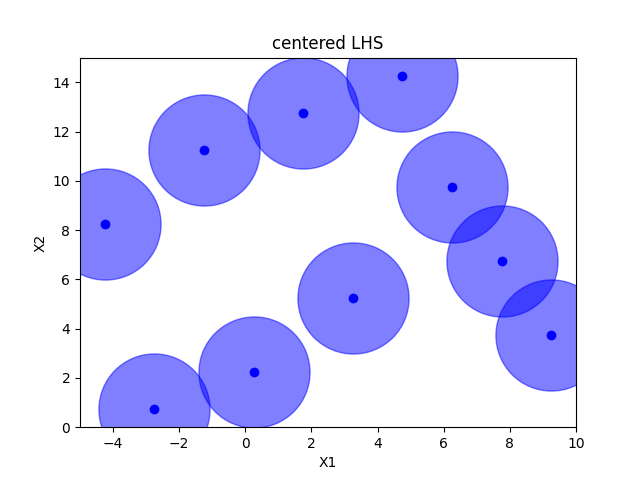
Centered Latin hypercube sampling#
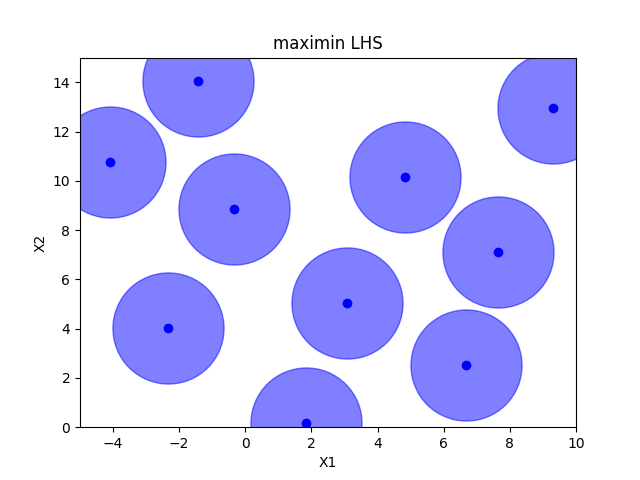
Maximin optimized hypercube sampling#
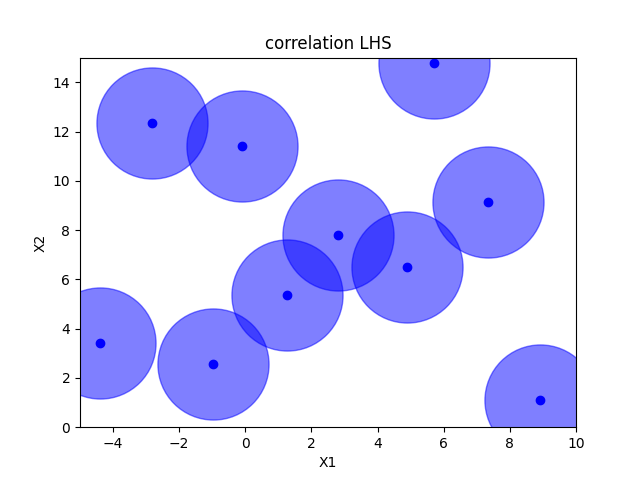
Correlation optimized hypercube sampling#
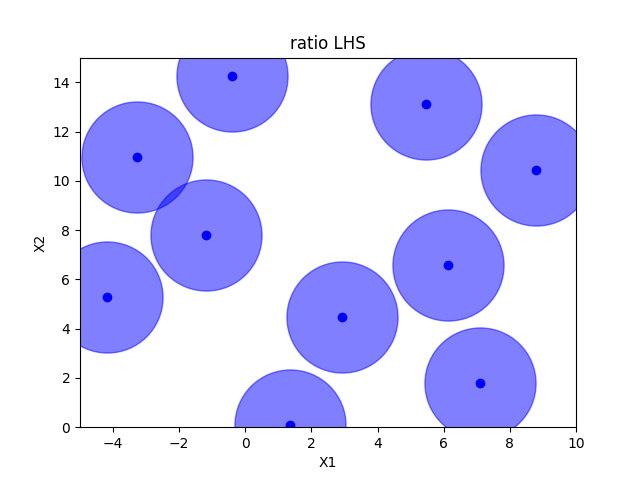
Ratio optimized hypercube sampling#
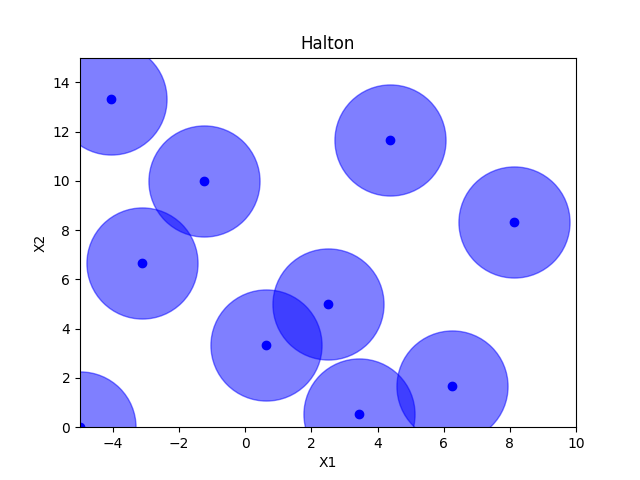
Halton sampling#
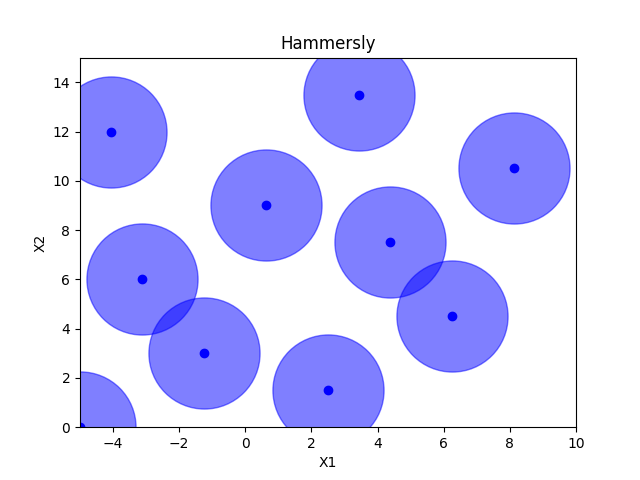
Hammersly sampling#
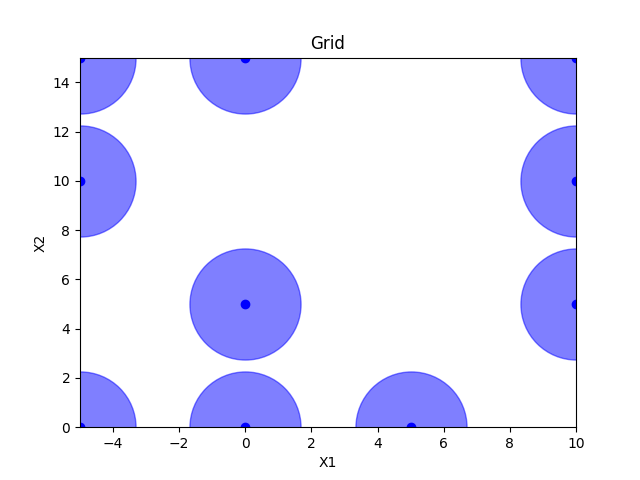
Grid sampling#
grid = Grid(border="include", use_full_layout=False)
x = grid.generate(space.dimensions, n_samples)
plot_searchspace(x, 'Grid')
pdist_data.append(pdist(x).flatten())
x_label.append("grid")
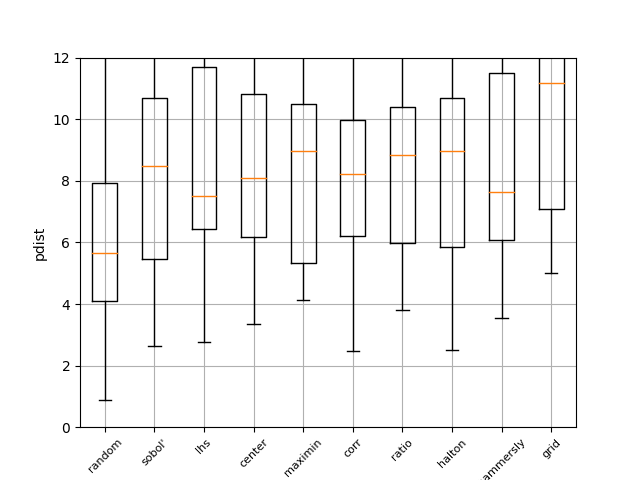
Pdist boxplot of all methods#
This boxplot shows the distance between all generated points using Euclidian distance. The higher the value, the better the sampling method. It can be seen that random has the worst performance
fig, ax = plt.subplots()
ax.boxplot(pdist_data)
plt.grid(True)
plt.ylabel("pdist")
_ = ax.set_ylim(0, 12)
_ = ax.set_xticklabels(x_label, rotation=45, fontsize=8)
Total running time of the script: (0 minutes 5.517 seconds)
